Insider Brief
- Researchers at Pusan National University have developed the Hypergraph Interactive Transformer (HIT), an AI model that significantly improves the accuracy and speed of identifying therapeutic gene targets using hypergraphs and attention-based learning.
- HIT outperformed existing models in predictive accuracy and computational efficiency, completing analysis in under two hours compared to weeks with traditional methods, while also enhancing explainability for medical researchers and clinicians.
- The AI model has the potential to revolutionize personalized medicine by accelerating drug discovery, improving early disease detection, and enabling treatments tailored to a patient’s genetic profile.
PRESS RELEASE -Traditional methods for identifying therapeutic gene targets, crucial for personalized medicine, are expensive and time-consuming. While artificial intelligence (AI), particularly deep learning approaches like deep graph representation learning, offer a promising alternative for identifying biomarker genes, they struggle to capture the complex, one-to-many relationships between diseases, genes, and gene ontologies, thus limiting their effectiveness in pinpointing therapeutic gene targets.
To address this challenge, a research team from Pusan National University, South Korea, led by Associate Professor Giltae Song from the School of Computer Science and Engineering, developed an innovative AI model called Hypergraph Interaction Transformer (HIT). “Our advanced AI model can not only predict gene-disease associations but also identify therapeutic gene targets with great precision. It utilizes hypergraph modelling and attention mechanisms that enable a comprehensive analysis of complex biological interactions,” explains Prof. Song. Their study was published in Volume 26, Issue 1, of the journal Briefings in Bioinformatics on January 22, 2025.
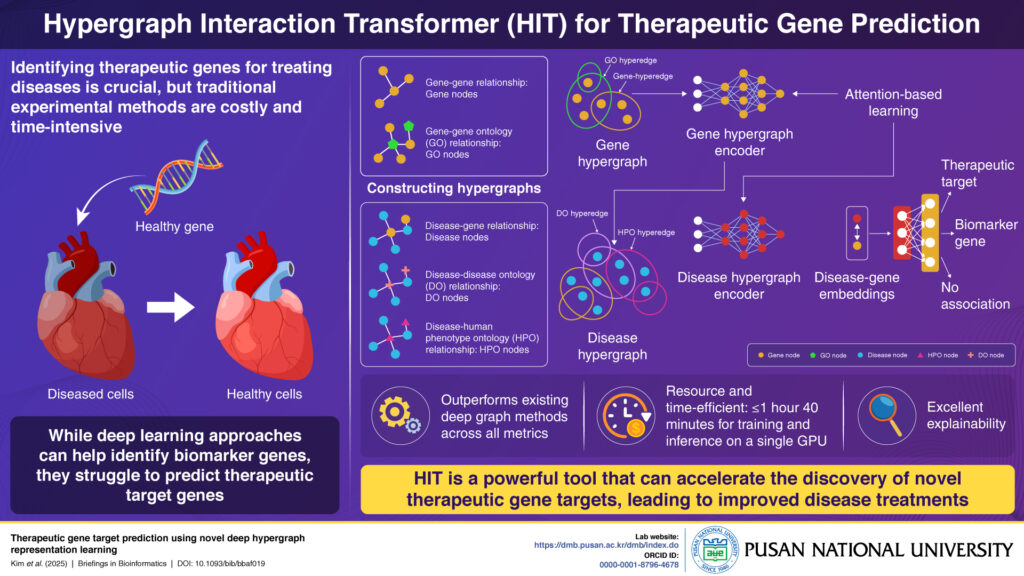
Proposed Hypergraph Interaction Transformer model (Image by Gitae Song from Pusan National University)
The HIT model utilizes hypergraphs, which, unlike traditional graphs, can connect multiple nodes with a single hyperedge. This allows HIT to effectively model complex biological relationships by constructing gene and disease hypergraphs from multiple biological datasets, capturing connections between genes, diseases, and various ontologies like gene, disease, and human phenotype ontologies.
Once the hypergraphs are constructed, the model processes them using two specialized encoders that use attention-based learning. The gene hypergraph encoder processes the gene hypergraph to create gene embeddings, which represent the relationship between a set of genes and the common gene ontology to which they are linked. These gene embeddings then serve as the initial embeddings for the corresponding genes in the disease hypergraph. The disease hypergraph encoder then refines the gene embeddings using the disease hypergraph and simultaneously produces new disease embeddings. Finally, the gene and disease embeddings are combined and used to specifically classify a gene as a therapeutic gene target, a biomarker, or unrelated to a specific disease.
HIT outperformed existing models in all tested metrics, demonstrating its accuracy in classifying therapeutic gene targets. Its efficiency is notable, requiring only 1 hour 40 minutes of single graphics processing unit-based training and inference, compared to weeks for traditional methods. A heart failure case study further validated its real-world applicability, successfully identifying all known therapeutic targets for the disease. Importantly, the model’s decision-making process is also highly explainable, allowing for increased trust from doctors and researchers.
“HIT can accelerate the discovery of novel therapeutic gene targets and contribute to the understanding of disease mechanisms,” notes Prof. Song. “This could advance personalized medicine by enabling treatments tailored to a patient’s genetic profile and improving early disease detection in clinical settings.”
By accurately and quickly identifying therapeutic gene targets, HIT can significantly shorten the drug development pipeline, allowing promising treatments to reach patients faster.
Reference
Title of original paper: Therapeutic gene target prediction using novel deep hypergraph representation learning
Journal: Briefings in Bioinformatics
DOI: 10.1093/bib/bbaf019


