Insider Brief
- A new benchmarking study found that some large language models (LLMs) can generate accurate, executable code for biomedical predictive modeling tasks in reproductive health, in some cases outperforming human-coded models.
- OpenAI’s o3-mini-high successfully completed 7 out of 8 tasks, with LLM-generated R code performing better overall than Python due to superior package support.
- While LLMs produced accurate models for predicting gestational age, their performance was slightly lower than human benchmarks in classifying preterm birth from microbiome data.
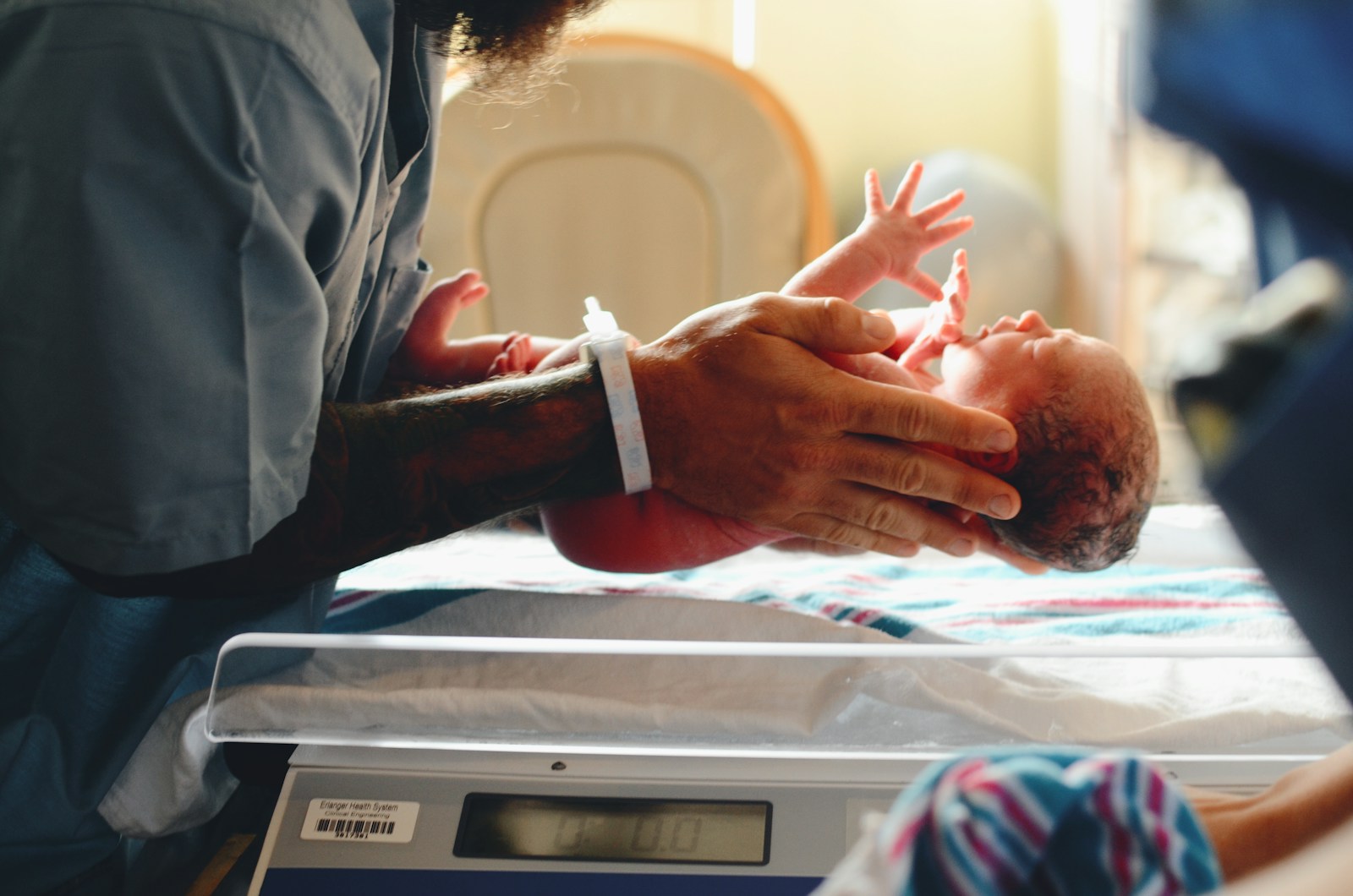
A new study has found that some large language models can match or even exceed human-coded models in predicting key outcomes in reproductive health, including gestational age and preterm birth risk.
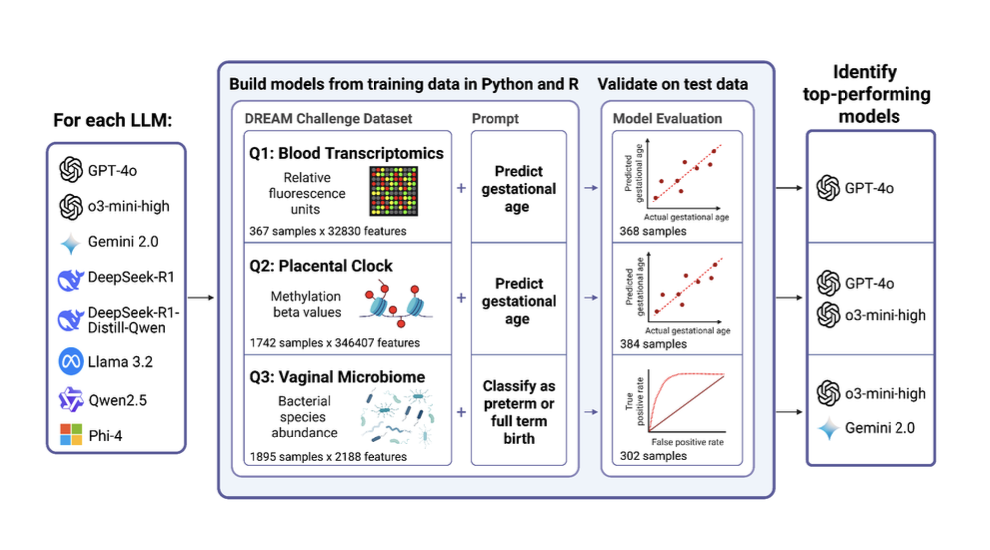
Published in the pre-print server bioRxiv, the study evaluated eight popular large language models (LLMs) on four predictive modeling tasks using biomedical datasets. The authors — spanning institutions from UC San Francisco to Wayne State and the NIH — tested whether these models could generate working R or Python code to analyze real-world omics data, produce usable predictions, and visualize results. The models were judged on their ability to complete the tasks successfully and the accuracy of their predictions on test datasets.
The research team used past DREAM Challenges as a testbed. These open science competitions crowdsource solutions to biomedical questions using anonymized, pre-partitioned datasets. In this case, the researchers selected tasks focused on reproductive health, including predicting gestational age from gene expression and DNA methylation data, and classifying preterm and early preterm birth based on microbiome profiles.
Of the LLMs tested, OpenAI’s o3-mini-high was the top performer, completing 7 out of 8 tasks successfully. Several other models, including 4o, DeepseekR1, and Gemini 2.0, completed at least one task without error. The models were assessed not only on their ability to produce functional code, but on whether the code returned accurate results when tested on blinded datasets from the original challenges.
Findings
The study found that LLM-generated R code outperformed Python code in task completion, with 14 out of 16 R-based analyses running successfully compared to just 7 Python-based runs. The team indicated that one key factor was the greater availability of robust R packages — particularly Bioconductor tools — that streamline access to public repositories like the Gene Expression Omnibus. On the other hand, LLMs struggled to write Python code capable of handling these data pulls correctly.
Despite those limitations, LLMs often performed competitively with or better than the human teams from the original DREAM Challenges. In one notable example, LLM-generated code for predicting gestational age from placental DNA methylation data produced a model with lower error than the top-scoring team in the original challenge. For that task, the model’s root mean squared error (RMSE) was 1.12 weeks, compared to 1.24 for the human-generated benchmark.
However, LLMs were less successful on classification tasks related to preterm birth. For example, predicting early preterm birth from microbiome data resulted in lower area under the ROC curve (AUROC) scores than the original human participants. Even so, several LLMs produced identical top-scoring models, raising questions about convergence and originality.
Clear Utility for LLMs
The authors argue the findings demonstrate a clear utility for LLMs in biomedical data science, particularly in fields like reproductive health where researchers may lack access to programming expertise. LLMs can automate routine data analysis tasks, generate reproducible workflows and potentially level the playing field across research groups with varying technical resources.
In this setting, LLMs were able to rapidly develop and run machine learning pipelines across multiple types of omics data — transcriptomic, methylation, and microbiome — without human intervention in the modeling process. While some manual post-processing was required (such as disabling automatic package installation or saving outputs), the LLMs largely succeeded in generating complete and runnable analysis scripts.
One of the motivations for this study was to address the “too much data, too few experts” bottleneck that plagues many areas of computational biology. By enabling non-coders to analyze complex datasets — or by helping coders move faster and more consistently — LLMs could help close gaps in biomedical research output, particularly for emerging areas like pregnancy risk stratification.
Methods
Each of the eight LLMs was prompted to generate code that performed three tasks for each dataset: fit a predictive model using training data, evaluate performance on a test set, and generate a relevant plot (either a scatter plot or ROC curve). The code was executed on a standardized computing platform with all required libraries pre-installed.
Scoring was based on four criteria: correct data extraction and formatting, successful model training and testing, accurate plotting, and top-tier performance compared to historical benchmarks. A maximum of five points could be awarded per task.
The highest scoring model, o3-mini-high, earned 33 out of a possible 40 points, and was the only model to generate complete and correct R code for all four tasks. In contrast, three models — DeepSeekDistill, Llama, and Phi-4 — failed to generate any usable code for any task.
Not a Panacea
The authors caution that LLMs are not a panacea. In many cases, multiple models generated identical outputs when given the same prompt, which could lead to reduced diversity in crowdsourced settings like DREAM Challenges. This homogeneity could blunt scientific insight by limiting the range of modeling strategies explored.
Another limitation was that the study relied on a single-shot prompt strategy — LLMs were not iteratively corrected or tuned. This reflects a more realistic use case for non-expert users but likely underestimates the models’ full potential if more interactive prompting were allowed.
Additionally, some of the test set advantages enjoyed by human challengers, such as access to supplemental data or feedback from early submissions, were not available to the LLMs, meaning the comparison may not be entirely apples-to-apples.
Next Steps
The researchers suggest that future studies could explore how fine-tuning LLMs on biomedical code repositories or clinical datasets might improve performance. More advanced prompt engineering strategies, model chaining, and feedback loops could also enhance usability and accuracy.
As the healthcare field moves toward greater reliance on machine learning and personalized prediction, tools like LLMs may become central in the pipeline—not as replacements for human expertise, but as accelerators for insight. Integrating these tools into clinical workflows will require rigorous validation, particularly when models influence high-stakes decisions like pregnancy management.
If you are looking for a deeper dive into this work, the research paper — available on bioRxiv — is quite technical and offers more information than this summary news article.
Pre-print servers, such as bioRxiv, are used by researchers to distribute their work in fast-changing fields, such as quantum science. However, these works have yet to be officially peer reviewed, a key step in the scientific method. Likewise, this article should not be considered a peer-review attempt.
The institutions represented in the study include the University of California, San Francisco; Huron High School in Ann Arbor, Michigan; New York University Langone Health; Wayne State University; the National Institutes of Health (specifically the Eunice Kennedy Shriver National Institute of Child Health and Human Development); the University of Michigan; and Michigan State University.